Balliu Lab @ UCLA
We are a newly minted (est. July 2022) computational research group in the Departments of Pathology, Computational Medicine, and Biostatistics at UCLA. We develop novel statistical methods and computational tools for analyzing high-dimensional repeated measures and intensive longitudinal data such as those arising from high-throughput genomic assays, mobile phone and wearable sensors, and electronic health records. We apply these methods to understand the genetic, molecular, cellular, and environmental mechanisms underlying complex human traits and diseases, with a focus on metabolic and psychiatric phenotypes.
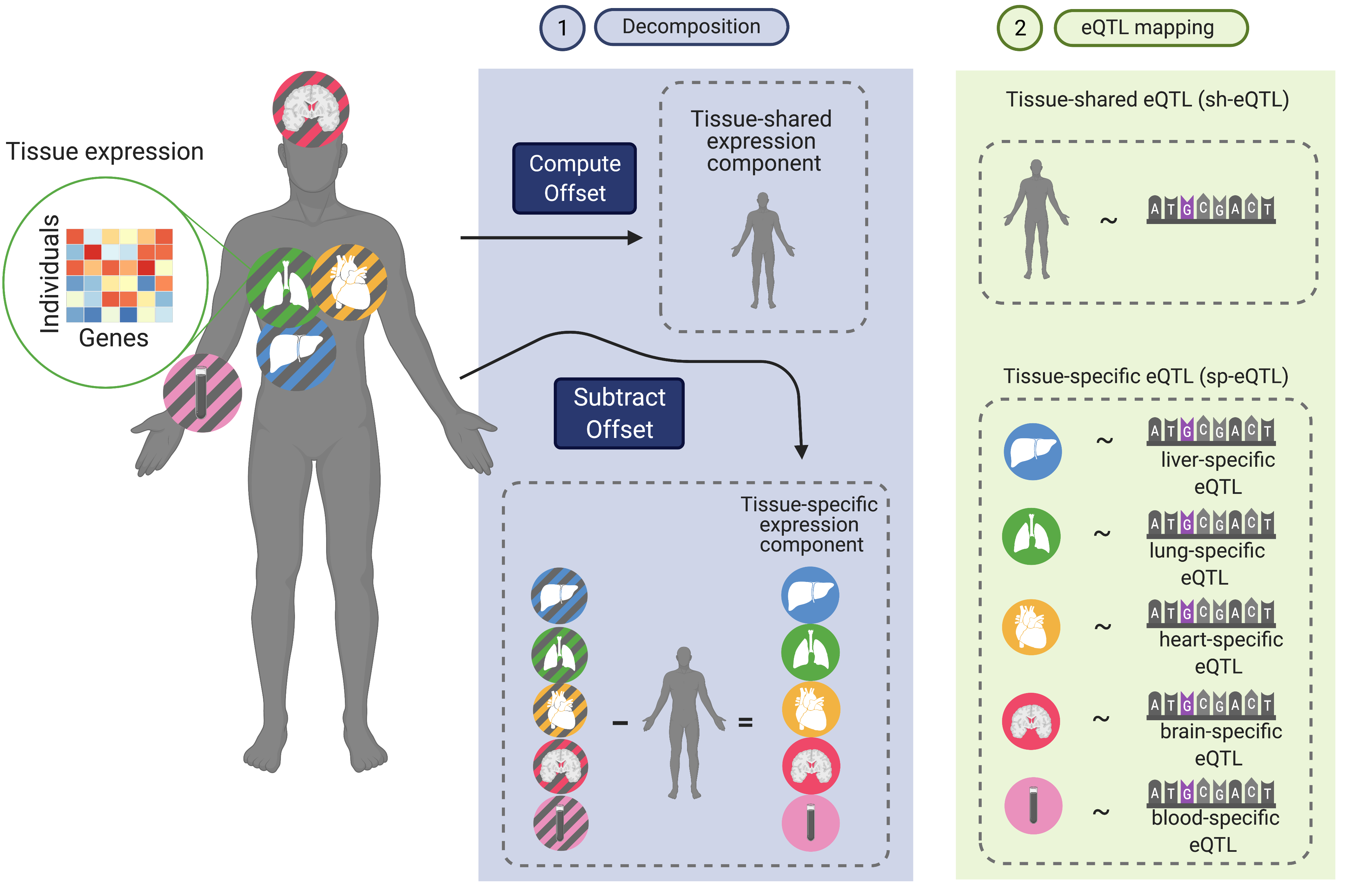
We are particularly interested in understanding the role of inherited genetic variation in disease through the diversity of changes observed in gene expression under different contexts, e.g. age, tissues & cell types, environmental perturbations, and reprogramming stages.
Join us!
Postdoctoral researchers
We have an opening for a postdoctoral position for quantitatively skilled PhDs to study the impact of genomic variation on the gene regulatory networks of reprogramming as part of the IGVF consortium. The position is shared with the Zaitlen lab at UCLA. If interested, please send me an e-mail with a short statement on your research interests, your CV, and contact information for three references.
Graduate students
We take UCLA students from Bioinformatics, Biomathematics, Biostatistics, Genetics and Genomics, Neuroscience and related programs. We mentor students with a strong foundation in quantitative disciplines (statistics, math, computer science, etc) who are interested in developing new methods. We also welcome students with diverse backgrounds, such as neuroscience or quantitative biology, who are interested in analyzing large-scale datasets with a focus on specific biological inquiries. If you believe you would be a good fit for the lab, please send me an e-mail with your research interests and your CV.
Undergraduate students
Unfortunately, we are not accepting undergrads this quarter. However, if you think you might be a good fit in the future, send me an e-mail with your research interests and your CV.